Photosynthesis is an electron transport process
The previous chapter described how photons are captured by an antenna and conducted as excitons to the reaction centers. This chapter deals with the function of these reaction centers and describes how photon energy is converted to chemical energy to be utilized by the cell. As mentioned in Chapter 2, plant photosynthesis probably evolved from bacterial pho- tosynthesis, so that the basic mechanisms of the photosynthetic reactions are alike in bacteria and plants. Bacteria have proved to be very suitable objects for studying the principles of photosynthesis since their reaction centers are more simply structured than those of plants and they are more easily isolated. For this reason, first bacterial photosynthesis and then plant photosynthesis will be described.
The photosynthetic machinery is constructed from modules
The photosynthetic machinery of bacteria is constructed from defined com- plexes, which also appear as components of the photosynthetic machinery in plants. As will be described in Chapter 5, some of these complexes are also components of mitochondrial electron transport. These complexes can be thought of as modules that developed at an early stage of evolu- tion and have been combined in various ways for different purposes. For easier understanding, the functions of these modules in photosynthesis will be treated first as black boxes and a detailed description of their structure and function will be given later.
E °
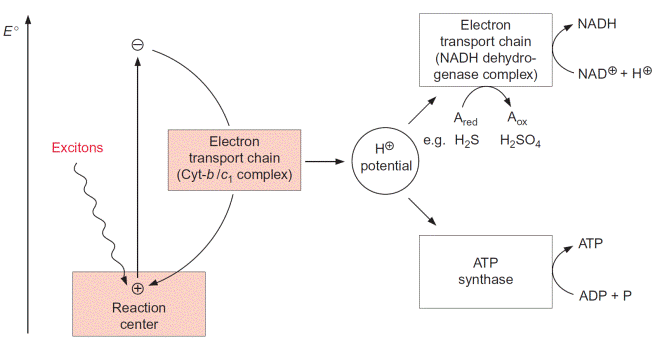
Figure 3.1 Schematic presentation of the photosynthetic apparatus of purple bacteria. The energy of a captured exciton in the reaction center elevates an electron to a negative redox state. The electron is transferred to the ground state via an electron transport chain including the cytochrome-b/c1 complex and cytochrome-c (the latter is not shown). Free energy of this process is conserved by formation of a proton potential which is used partly for synthesis of ATP and partly to enable an electron flow for the formation of NADH from electron donors such as H2S.
========================================================
Oxidation state shows the total number of electrons which have been removed from an element (a positive oxidation state) or added to an element (a negative oxidation state) to get to its present state.
- Oxidation involves an increase in oxidation state
- Reduction involves a decrease in oxidation state
Section 6.1 – An Introduction to Oxidation Reduction Reactions (10:27)
Section 6.2 – Oxidation Numbers (22:09)
Section 6.4 – Voltaic Cells (24:17)
0.3: Electrochemical Potential
n a galvanic cell, current is produced when electrons flow externally through the circuit from the anode to the cathode because of a difference in potential energy between the two electrodes in the electrochemical cell. In the Zn/Cu system, the valence electrons in zinc have a substantially higher potential energy than the valence electrons in copper because of shielding of the s electrons of zinc by the electrons in filled d orbitals. Hence electrons flow spontaneously from zinc to copper(II) ions, forming zinc(II) ions and metallic copper. Just like water flowing spontaneously downhill, which can be made to do work by forcing a waterwheel, the flow of electrons from a higher potential energy to a lower one can also be harnessed to perform work….
Redox reactions can be balanced using the half-reaction method. The standard cell potential is a measure of the driving force for the reaction. \(E°_{cell} = E°_{cathode} − E°_{anode} \] The flow of electrons in an electrochemical cell depends on the identity of the reacting substances, the difference in the potential energy of their valence electrons, and their concentrations. The potential of the cell under standard conditions (1 M for solutions, 1 atm for gases, pure solids or liquids for other substances) and at a fixed temperature (25°C) is called the standard cell potential (E°cell). Only the difference between the potentials of two electrodes can be measured. The potential of the standard hydrogen electrode (SHE) is defined as 0 V under standard conditions. The potential of a half-reaction measured against the SHE under standard conditions is called its standard electrode potential. By convention, all tabulated values of standard electrode potentials are listed as standard reduction potentials. The overall cell potential is the reduction potential of the reductive half-reaction minus the reduction potential of the oxidative half-reaction (E°cell = E°cathode − E°anode). The standard cell potential is a measure of the driving force for a given redox reaction. If E°cell is positive, the reaction will occur spontaneously under standard conditions. If E°cell is negative, then the reaction is not spontaneous under standard conditions, although it will proceed spontaneously in the opposite direction.
Cyclic Electron Transport in Photosynthesis
Photosynthetic reaction center
The reaction center is in the thylakoid membrane. It transfers light energy to a dimer of chlorophyll pigment molecules near the periplasmic (or thylakoid lumen) side of the membrane. This dimer is called a special pair because of its fundamental role in photosynthesis. This special pair is slightly different in PSI and PSII reaction center. In PSII, it absorbs photons with a wavelength of 680 nm, and it is therefore called P680. In PSI, it absorbs photons at 700 nm, and it is called P700. In bacteria, the special pair is called P760, P840, P870, or P960. “P” here means pigment, and the number following it is the wavelength of light absorbed.
If an electron of the special pair in the reaction center becomes excited, it cannot transfer this energy to another pigment using resonance energy transfer. In normal circumstances, the electron should return to the ground state, but, because the reaction center is arranged so that a suitable electron acceptor is nearby, the excited electron can move from the initial molecule to the acceptor. This process results in the formation of a positive charge on the special pair (due to the loss of an electron) and a negative charge on the acceptor and is, hence, referred to as photoinduced charge separation. In other words, electrons in pigment molecules can exist at specific energy levels. Under normal circumstances, they exist at the lowest possible energy level they can. However, if there is enough energy to move them into the next energy level, they can absorb that energy and occupy that higher energy level. The light they absorb contains the necessary amount of energy needed to push them into the next level. Any light that does not have enough or has too much energy cannot be absorbed and is reflected. The electron in the higher energy level, however, does not want to be there; the electron is unstable and must return to its normal lower energy level. To do this, it must release the energy that has put it into the higher energy state to begin with. This can happen various ways. The extra energy can be converted into molecular motion and lost as heat. Some of the extra energy can be lost as heat energy, while the rest is lost as light. (This re-emission of light energy is called fluorescence.) The energy, but not the e- itself, can be passed onto another molecule. (This is called resonance.) The energy and the e- can be transferred to another molecule. Plant pigments usually utilize the last two of these reactions to convert the sun’s energy into their own.
This initial charge separation occurs in less than 10 picoseconds (10−11 seconds). In their high-energy states, the special pigment and the acceptor could undergo charge recombination; that is, the electron on the acceptor could move back to neutralize the positive charge on the special pair. Its return to the special pair would waste a valuable high-energy electron and simply convert the absorbed light energy into heat. In the case of PSII, this backflow of electrons can produce reactive oxygen species leading to photoinhibition.[1][2] Three factors in the structure of the reaction center work together to suppress charge recombination nearly completely.
- Another electron acceptor is less than 10 Å away from the first acceptor, and so the electron is rapidly transferred farther away from the special pair.
- An electron donor is less than 10 Å away from the special pair, and so the positive charge is neutralized by the transfer of another electron
- The electron transfer back from the electron acceptor to the positively charged special pair is especially slow. The rate of an electron transfer reaction increases with its thermodynamic favorability up to a point and then decreases. The back transfer is so favourable that it takes place in the inverted region where electron-transfer rates become slower.[1]
Thus, electron transfer proceeds efficiently from the first electron acceptor to the next, creating an electron transport chain that ends if it has reached NADPH.
========================================================
Purple bacteria have only one reaction center (Fig. 3.1). In this reac- tion center the energy of the absorbed photon excites an electron, which will be elevated to a negative redox state. The excited electron is transferred back to the ground state by an electron transport chain, called the cyto- chrome-b/c1 complex, and the released energy is transformed to a chemical compound (NADH), which is then used for the synthesis of biomass (e.g., proteins and carbohydrates). Generation of energy is based on coupling the electron transport with the transport of protons across the membrane. In this way the energy of the excited electron is conserved as an electrochemical H-potential across the membrane. The photosynthetic reaction cent- ers and the main components of the electron transport chain are always located in a membrane.
Via ATP-synthase the energy of the H-potential is used to synthesize ATP from ADP and phosphate. Since the excited electrons in purple bacte- ria return to the ground state of the reaction center, this electron transport is called cyclic electron transport. In purple bacteria the proton gradient is also used to reduce NAD via an additional electron transport chain named the NADH dehydrogenase complex (Fig. 3.1). By consuming the energy of the H-potential, electrons are transferred from a reduced compound (e.g., organic acids or hydrogen sulfide) to NAD. The ATP and NADH formed by bacterial photosynthesis are used for the synthesis of organic matter; especially important is the synthesis of carbohydrates from CO2 via the Calvin cycle (see Chapter 6).
The reaction center of green sulfur bacteria (Fig. 3.2) is homologous to that of purple bacteria, indicating that they have both evolved from a com- mon ancestor. ATP is also formed in green sulfur bacteria by cyclic electron transport. The electron transport chain (cytochrome-b/c1 complex) and the ATP-synthase involved here are very similar to those in purple bacteria. However, in contrast to purple bacteria, green sulfur bacteria are able to synthesize NADH by a noncyclic electron transport process. In this case, the excited electrons are transferred to the ferredoxin-NAD-reductase complex, which reduces NAD to NADH. Since the excited electrons in this noncyc- lic pathway do not return to the ground state, an electron deficit remains in the reaction center and is replenished by electron donors such as H2S, ulti- mately being oxidized to sulfate.
Cyanobacteria and plants use water as an electron donor in photosyn thesis (Fig. 3.3). As oxygen is liberated, this process is called oxygenic pho- tosynthesis. Two photosystems designated II and I are arranged in tandem. The machinery of oxygenic photosynthesis is built by modules that have already been described in bacterial photosynthesis. The structure of the reaction center of photosystem II corresponds to that of the reaction center of purple bacteria, and that of photosystem I corresponds to the reaction center of green sulfur bacteria. The enzymes ATP synthase and ferredoxin- NADP-reductase are very similar to those of photosynthetic bacteria. The
Figure 3.2 Schematic presentation of the photosynthetic apparatus in green sulfur bacteria. In contrast to the scheme in Figure 3.1, part of the electrons elevated to a negative redox state is transferred via an electron transport chain (ferredoxin- NAD reductase) to NAD, yielding NADH. The electron deficit arising in the reaction center is compensated by electron donors such as H2S (see also Fig. 3.1).
Figure 3.3 Schematic presentation of the photosynthetic apparatus of cyanobacteria and plants. The two sequentially arranged reaction centers correspond in their function to the photosynthetic reaction centers of purple bacteria and green sulfur bacteria (shown in Figs. 3.1 and 3.2).
electron transport chain of the cytochrome-b6/f complex has the same basic structure as the cytochrome-b/c1 complex in bacteria.
Four excitons are required in oxygenic photosynthesis to split one mol- ecule of water:
H2O NADP 4 excitons → 1/2 O2 NADPH H
In this noncyclic electron transport, electrons are transferred to NADP and protons are transported across the membrane to generate a proton gradient that drives the synthesis of ATP. Thus, for each mol of NADPH formed by oxygenic photosynthesis, about 1.5 molecules of ATP are gener- ated simultaneously (section 4.4). Most of this ATP and NADPH are used for CO2 and nitrate assimilation to synthesize carbohydrates and amino acids. Oxygenic photosynthesis in plants takes place in the chloroplasts, a cell organelle of the plastid family (section 1.3).
Vides
Redox Reactions
o-Oxidation and reduction review from biological point-of-view | Biomolecules | MCAT | Khan Academy
Oxidation and reduction in cellular respiration | Biology | Khan Academy
A reductant and an oxidant are formed during photosynthesis
In the 1920s Otto Warburg (Berlin) postulated that the energy of light is transferred to CO2 and that the CO2, activated in this way, reacts with water to form a carbohydrate, accompanied by the release of oxygen. According to this hypothesis, the oxygen released by photosynthesis was derived from the CO2. In 1931 this hypothesis was opposed by Cornelis van Niel (USA) by postulating that during photosynthesis a reductant is formed, which then reacts with CO2. The so-called van Niel equation describes photosynthesis in the following way: He proposed that a compound H2A is split by light energy into a reduc- ing compound (2H) and an oxidizing compound (A). For oxygenic photo- synthesis of cyanobacteria or plants, it can be rewritten as:

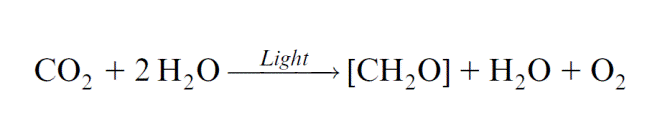
In this equation the oxygen released during photosynthesis is derived from water. In 1937 Robert Hill (Cambridge, UK) proved that a reductant is actu- ally formed in the course of photosynthesis. He was the first to succeed in isolating chloroplasts with photosynthetic activity, which, however, had no intact envelope membranes and consisted only of thylakoid membranes. When these chloroplasts were illuminated in the presence of Fe3 com- pounds (initially ferrioxalate, later ferricyanide ([Fe (CN)6]3)), Robert Hill observed an evolution of oxygen accompanied by the reduction of the Fe3-compounds to the Fe2 form.

This “Hill reaction” proved that the photochemical splitting of water can be separated from the reduction of the CO2. Therefore the complete reac- tion of photosynthetic CO2 assimilation can be divided into two reactions:
- The so-called light reaction, in which water is split by photon energy to yield reductive power (NADPH) and chemical energy (ATP); and
- the so-called dark reaction (Chapter 6), in which CO2 is assimilated at the expense of the reductive power and of ATP.
In 1952 the Dutchman Louis Duysens made a very important observa- tion that helped explain the mechanism of photosynthesis. When illumi- nating isolated membranes of the purple bacterium Rhodospirillum rubrum with short light pulses, he found a decrease in light absorption at 890 nm, which was immediately reversed when the bacteria were darkened again. The same “bleaching” effect was found at 870 nm in the purple bacte- rium Rhodobacter sphaeroides. Later, Bessil Kok (USA) and Horst Witt (Germany) also found similar pigment bleaching at 700 nm and 680 nm in chloroplasts. This bleaching was attributed to the primary reaction of pho- tosynthesis, and the corresponding pigments of the reaction centers were named P870 (Rb. sphaeroides) and P680 and P700 (chloroplasts). When an oxidant (e.g., [Fe(CN)6]3) was added, this bleaching effect could also be achieved in the dark. These results indicated that these absorption changes of the pigments were due to a redox reaction. This was the first indication that chlorophyll can be oxidized. Electron spin resonance measurements revealed that radicals are formed during this “bleaching.” “Bleaching” could also be observed at the very low temperature of 1 K. This showed that in the electron transfer leading to the formation of radicals, the reac- tion partners are located so close to each other that thermal oscillation of the reaction partners (normally the precondition for a chemical reaction) is not required for this redox reaction. Spectroscopic measurements indicated that the reaction partner of this primary redox reaction are two closely adjacent chlorophyll molecules arranged as a pair, called a “special pair.”
The basic structure of a photosynthetic reaction center has been resolved by X-ray structure analysis
The reaction centers of purple bacteria proved to be especially suitable objects for explaining the structure and function of the photosynthetic machinery. It was a great step forward when in 1970 Roderick Clayton (USA) developed a method for isolating reaction centers from purple bacte- ria. Analysis of the components of the reaction centers of the different purple bacteria (shown in Table 3.1 for the reaction center of Rhodobacter sphaer- oides as an example) revealed that the reaction centers had the same basic structure in all the purple bacteria investigated. The minimum structure
consists of the three subunits L, M, and H (light, medium, and heavy). Subunits L and M are peptides with a similar amino acid sequence. They are homologous. The reaction center of Rb. sphaeroides contains four bac- teriochlorophyll-a (Bchl-a, Fig. 3.4) and two bacteriopheophytin-a (Bphe- a). Pheophytins differ from chlorophylls in that they lack magnesium as the central atom. In addition, the reaction center contains an iron atom that is not part of a heme. It is therefore called a non-heme iron. Furthermore, the reaction center is comprised of two molecules of ubiquinone (Fig. 3.5), which are designated as QA and QB. QA is tightly bound to the reaction center, whereas QB is only loosely associated with it.
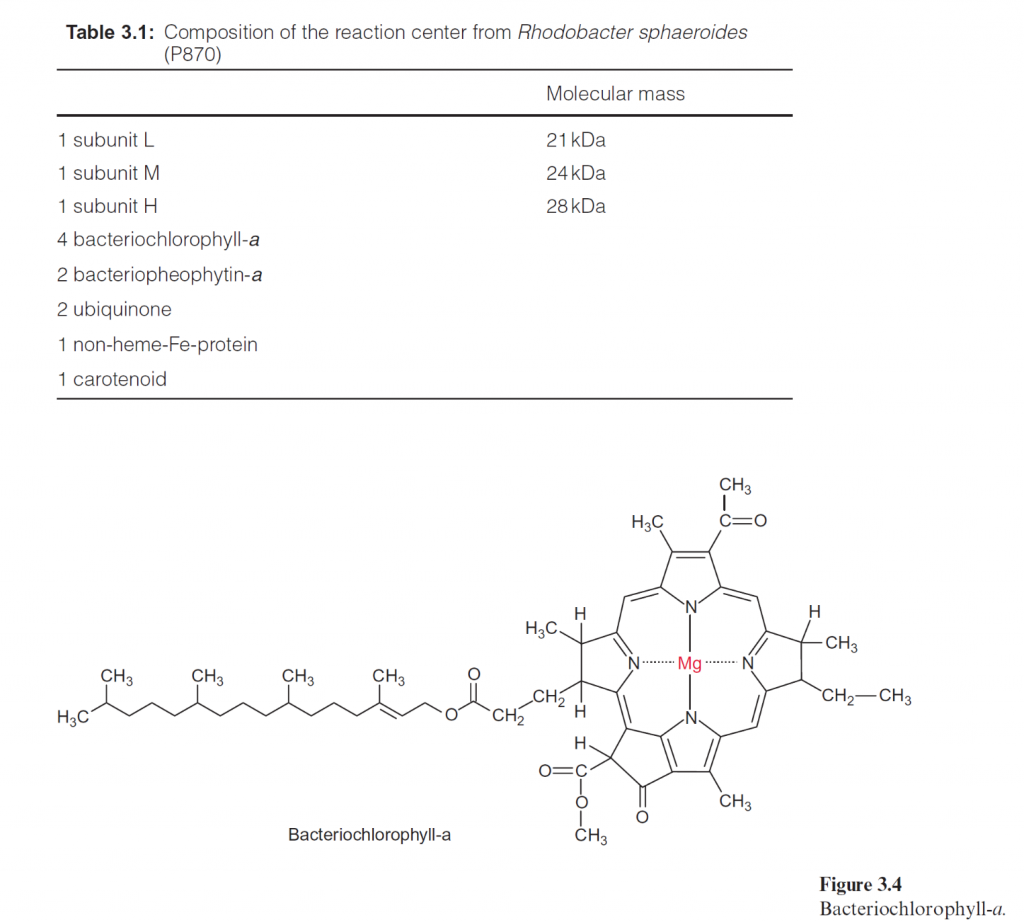
Figure 3.5 Ubiquinone. The long isoprenoid side chain mediates the lipophilic character and membrane anchorage.
X-ray structure analysis of the photosynthetic reaction center
If ordered crystals can be prepared from a protein, it is possible to analyze the spherical structure of the protein molecule by X-ray structure analysis. In this method a protein crystal is exposed to X-ray irradiation. The elec- trons of the atoms in the molecule cause a scattering of X-rays. Diffraction is observed when the irradiation passes through a regular repeating struc- ture. The corresponding diffraction pattern, consisting of many single reflections, is measured by an X-ray film positioned behind the crystal or by an alternative detector. The principle is demonstrated in Figure 3.6. To obtain as many reflections as possible, the crystal, mounted in a capillary, is rotated. From a few dozen to up to several hundred exposures are required for one set of data, depending on the form of the crystal and the size of the crystal lattice. To evaluate a new protein structure, several sets of data are required in which the protein has been changed by the incorporation or binding of a heavy metal ion. With the help of elaborate computer pro- grams, it is possible to reconstruct the spherical structure of the exposed protein molecules by applying the rules for scattering X-rays by atoms of various electron densities. slowly and the diffraction pattern is monitored on an X-ray film. Nowadays much more sensitive detector systems (image platers) are used instead of films. The diffraction pattern shown was obtained by the structural analysis of the reaction center of
Rb. sphaeroides. (Courtesy of H. Michel, Frankfurt.)
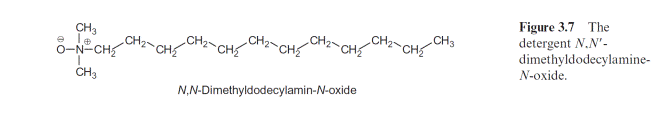
X-ray structure analysis requires a highly technical expenditure and is very time-consuming, but the actual limiting factor in the elucidation of a spherical structure is usually the preparation of suitable single crystals. Until 1980 it was thought to be impossible to prepare crystals suitable for X-ray structure analysis from hydrophobic membrane proteins. The appli- cation of the detergent N,N-dimethyldodecylamine-N-oxide (Fig. 3.7) was a great step forward in helping to solve this problem. This detergent forms water-soluble protein-detergent micelles with membrane proteins. With the addition of ammonium sulfate or polyethylene glycol is was then pos- sible to crystallize membrane proteins. The micelles form a regular lattice in these crystals (Fig. 3.8). The protein in the crystal remains in its native state since the hydrophobic regions of the membrane protein, which nor- mally border on the hydrophobic membrane, are covered by the hydropho- bic chains of the detergent.
Using this procedure, Hartmut Michel (Munich) succeeded in obtaining crystals from the reaction center of the purple bacterium Rhodopseudomonas viridis and, together with his colleagues Johann Deisenhofer and Robert Huber, performed an X-ray structure analysis of these crystals. The immense amount of time invested in these investigations is illustrated by the fact that the evaluation of the stored data sets alone took two and a half years (nowa- days modern computer programs would do it very much faster). The X-ray structure analysis of a photosynthetic reaction center successfully elucidated for the first time the three-dimensional structure of a membrane protein. For this work, Michel, Deisenhofer, and Huber were awarded the Nobel Prize in Chemistry in 1988. Using the same method, the reaction center of Rb. sphaeroides was analyzed and it turned out that the basic structures of the two reaction centers are astonishingly similar.

Figure 3.6 Schematic presentation of X-ray structural analysis of a protein crystal. A capillary containing the crystal rotates slowly and the diffraction pattern is monitored on an X-ray film. Nowadays muc more sensitive detector systems (image platers) are used instead of films. The diffraction pattern shown was obtained by the structural analysis of the reaction center of Rb. sphaeroides. (Courtesy of H. Michel, Frankfurt.)
The reaction center of Rhodopseudomonas viridis has a symmetric structure
Figure 3.9 shows the three-dimensional structure of the reaction center of the purple bacterium Rhodopseudomonas viridis. The molecule has a cylin- drical shape and is about 8 nm long. The homologous subunits L (red) and

Figure 3.8 A micelle is formed after solubilization of a membrane protein with detergent. The hydrophobic region of the membrane proteins, the membrane lipids, and the detergent are shown in black and the hydrophilic regions in red. Crystal structures can be formed by association of the hydrophilic regions of the detergent micelle.
M (black) are arranged symmetrically and enclose the chlorophyll and phe- ophytin molecules. The H subunit is attached like a lid to the lower part of the cylinder. In the same projection as in Figure 3.9, Figure 3.10 shows the location of the chromophores in the protein molecule. All the chromophores are positioned as pairs divided by a symmetry axis. Two Bchl-a molecules (DM, DL) can be recognized in the upper part of the structure. The two tetrapy- rrole rings are so close (0.3 nm) that their orbitals overlap in the excited state. This confirmed the actual existence of the “special pair” of chloro- phyll molecules, postulated earlier from spectroscopic investigations, as the
site of the primary redox process of photosynthesis. The chromophores are arranged underneath the chlorophyll pair in two nearly identical branches, both comprised of a Bchl-a (BA, BB) monomer and a bacteriopheophytin (A, B). Whereas the chlorophyll pair (DM, DL) is bound by both sub- units L and M, the chlorophyll BA and the pheophytin A are associated with subunit L, and BB and B with subunit M. The quinone ring of QA is bound via hydrogen bonds and hydrophobic interaction to subunit M, whereas the loosely associated QB is bound to subunit L.
playlist–
Photosynthesis
Photosystem II Function: The P680 Reaction Center
How does a reaction center function?
The analysis of the structure and extensive kinetic investigations allowed a detailed description of the function of the bacterial reaction center. The kinetic investigations included measurements by absorption and fluores- cence spectroscopy after light flashes in the range of less than 1013 s, as well as measurements of nuclear spin and electron spin resonance. Although the reaction center shows a symmetry with two almost identical branches of chromophores, electron transfer proceeds only along the right branch (the L side, Fig. 3.10). The chlorophyll monomer (BB) on the M side is in close contact with a carotenoid molecule, which abolishes a harmful triplet state of chlorophylls in the reaction center (sections 2.3 and 3.7). The function of the pheophytin (B) on the M side and of the non-heme iron is not yet fully understood.
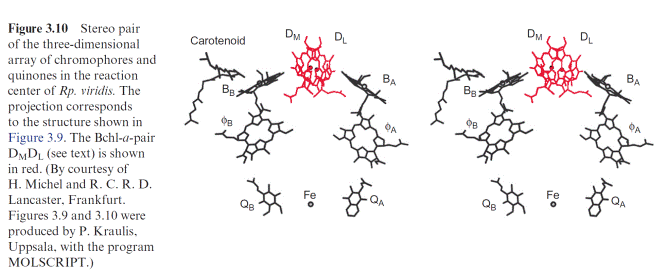
Figure 3.11 presents a scheme of the reaction center with the reaction partners arranged according to their electrochemical potential. The exciton of the primary reaction is provided by the antenna (section 2.4) which then excites the chlorophyll pair. This primary excitation state has only a very short half-life time, then a charge separation occurs, and, as a result of the large potential difference, an electron is removed within picoseconds to reduce bacteriopheophytin (Bphe).

The electron is probably transferred first to the Bchl-monomer (BA) and then to the pheophytin molecule (A). The second electron transfer proceeds with a half-time of 0.9 picoseconds, about four times as fast as the elec- tron transfer to BA. The pheophytin radical has a tendency to return to the ground state by a return of the translocated electron to the Bchl-monomer (BA). To prevent this, within 200 picoseconds a high potential difference withdraws the electron from the pheophytin radical to a quinone (QA) (Fig. 3.11). The semiquinone radical thus formed, in response to a further poten- tial difference, transfers its electron to the loosely bound ubiquinone QB. After a second electron transfer, first ubisemiquinone and then ubihydroqui- none are formed (Fig. 3.12). In contrast to the very labile radical intermedi- ates of the pathway, ubihydroquinone is a stable reductant. However, this stability has its price. For the formation of ubihydroquinone as a first sta- ble product from the primary excitation state of the chlorophyll, more than half of the exciton energy is dissipated as heat.
Ubiquinone (Fig. 3.5) contains a hydrophobic isoprenoid side chain, by which it is soluble in the lipid phase of the photosynthetic membrane. The same function of an isoprenoid side chain has already been discussed in the case of chlorophyll (section 2.2). In contrast to chlorophyll, pheophytin, and QA, which are all tightly bound to proteins, ubihydroquinone QB is only loosely associated with the reaction center and can be exchanged by another ubiquinone. Ubihydroquinone remains in the membrane phase, is able to diffuse rapidly along the membrane, and functions as a transport metabolite for reducing equivalents in the membrane phase. It feeds the electrons into the cytochrome-b/c1 complex, also located in the membrane. The electrons are then transferred back to the reaction center through the cytochrome-b/c1
complex and via cytochrome-c. Energy is conserved during this electron transport as a proton potential (section 4.1), which is used for ATP-synthe- sis. The structure and mechanism of the cytochrome-b/c1 complex and of ATP-synthase will be described in section 3.7 and Chapter 4, respectively.
In summary, the cyclic electron transport of the purple bacteria resembles a simple electric circuit (Fig. 3.13). The two pairs of chlorophyll and pheophy- tin, between which an electron is transferred by light energy, may be regarded as the two plates of a capacitor between which a voltage is generated, driv- ing a flux of electrons, a current. Voltage drops via a resistor and a large amount of the electron energy is dissipated as heat. This resistor functions
as an electron trap, and withdraws the electrons rapidly from the capacitor. A generator utilizes the remaining voltage to produce chemical energy.

Figure 3.11 Schematic presentation of cyclic electron transport in photosynthesis of Rb. sphaeroides. The excited state symbolized by a star results in a charge separation; an electron is transferred via pheophytin, the quinones QA, QB, and the cyt-b/c complex to the positively charged chlorophyll radical. Q: quinone, Q• : semiquinone radical, QH2: hydroquinone.

Figure 3.12 Reduction ofa quinone by one electronresults in a semiquinoneradical and furtherreduction to hydroquinone.Q: quinone, Q• :semiquinone radical, QH2:hydroquinone.ExcitonGeneratorCyt-b/c1complexElectrontrap+ –Chlorophyll dimerHeatchemicalworkATP
Figure 3.13 Cyclic electrontransport of photosynthesisdrawn as an electricalcircuit.
